 RAD51 D187 residue is displayed in green. BRC4 T1526A is a previously characterized missense variant that disrupts RAD51 binding. Bottom Insets: predicted disruptions resulting from variants BRC2 (S1221P), BRC4 (T1526A), and BRC7 (T1980I). Top Inset: BRC2 (S1221), BRC4 (T1526), and BRC7 (T1980) overlayed to show conservation. Homology model comparison of BRC2 (orange) and BRC7 (yellow) based on the BRC4 (pink) structure. ( C) Structural models based on 1N0W structure. FxTASGK is a consensus motif important for RAD51 interactions depicted below BRC2 and BRC7. Uniprot and ClustalX (70% threshold for shading) were used for the alignment. 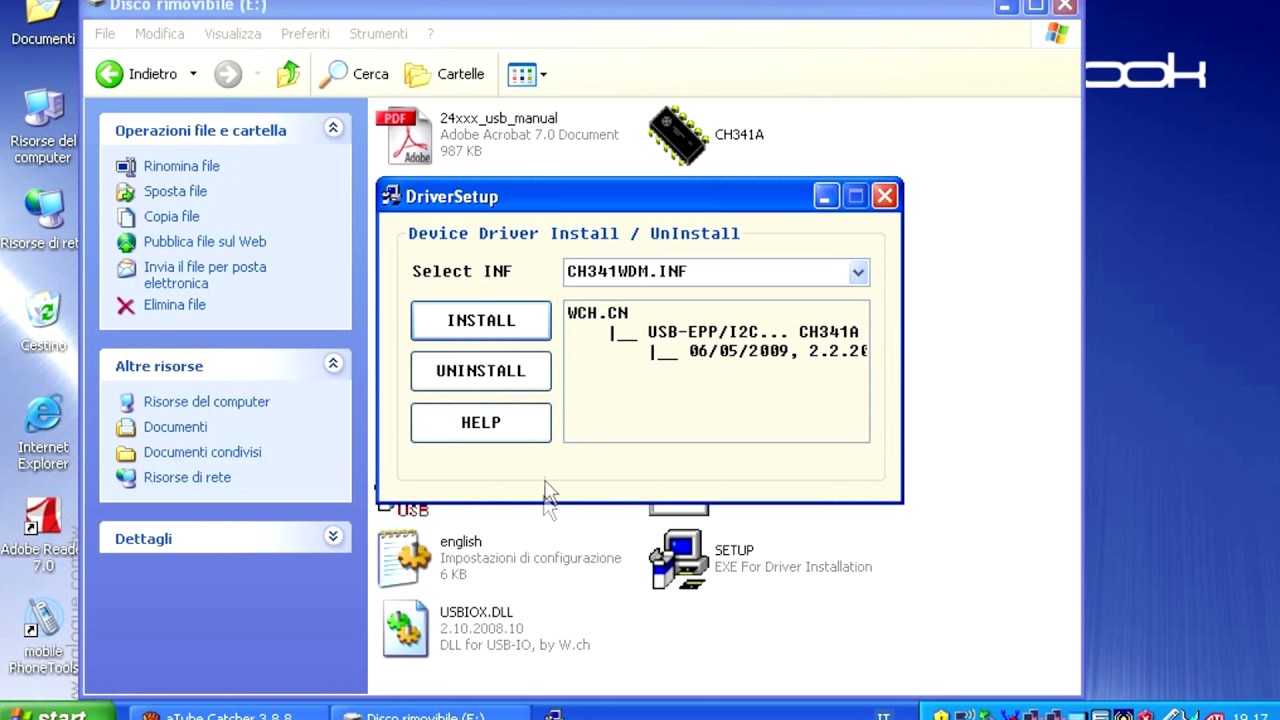
Residue corresponding to the missense variant is indicated in pink and conserved residues flanking the missense residue are indicated in light grey. ( B) Multiple sequence alignment of BRCA2 amino acids flanking each missense variant from different organisms: S1221P (BRC2), T1346I (Spacer BRC2-3), and T1980I (BRC7). 
BRCA2 missense variants used in this study are indicated. 
( A) BRCA2 protein schematic depicting domain organization: 2XMBP tag, N-terminus, BRC repeats, DNA binding domain (DBD), and C-terminal domain (CTD). S1221P and T1980I are structurally predicted to disrupt BRC folding and RAD51 binding. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
